In a recent
paper, Metz et al. describe a statistical framework for constructing the kinome polypharmacology network. Using a large
kinase panel (i.e., more than 150,000 activity values across 172 kinases for 3,858 compounds), they propose a set of statistical parameters (based on some measures of reliability and relevance) that can be used to establish relationships between kinases. Although such networks are useful in analyzing polypharmacology trends, the authors subsequently provide the following insight:
In order to impact medicinal chemistry design decisions, what is required is an understanding of specific chemotypes or functional groups that either create or destroy a specific connection, as defined by the therapeutic endpoint.
To that end, we've recently put together a tool, the
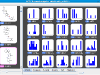
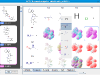
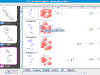
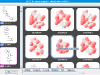
kinome navigator, that provides chemotype-centric views of kinase data. It's based on an improved version of our automated R-group analysis tool. The input requirement is the same as that of the kinome viewer, namely, kinase activity values are expressed in nM. Here is the
proper input of the above dataset (which is also bundled with the tool). Below are some sample screenshots.
Click on the button below to launch it. Please note that the tool is memory intensive, so it's best to run it on a machine with at least 1Gb of memory. As always, we welcome comments and/or suggestions.