Automated R-group analysis
Update Apr. 27, 2013: This tool has been obsoleted by Scaffold Hopper with additional features and bug fixes.
Generating R-group tables is a tedious task that we often find ourselves performed repeatedly for every project. For a typical HTS campaign, this task can be as simple as taking the most potent compounds and performing substructure searches around the identified scaffolds. For most cases, however, the task is quite involved, requiring a combination of clustering, substructure-, and pair-wise MCS searches. Here a couple of issues with this approach should be noted. First, clustering results are very sensitive to not only the underlying algorithm, but also on the metric used to compute the distance/similarity values between molecular fingerprints. We have seen commercial molecular clustering software failed repeatedly on fairly obvious cases. And secondly, it doesn't take much imagination to come up with examples where scaffolds derived from maximum common substructure searches are clearly not the "correct" scaffolds. For this very reason we're often puzzled as to why most cheminformatics toolkits (we're only aware of a couple of exceptions) don't provide an (exact) implementation for generating/enumerating all maximal common substructures.
Recently, we developed a prototype that can automatically generate R-group tables for any reasonable sized collection (limited by the available CPU cores and physical memory) of med-chem friendly compounds. Our implementation---in contrast to the one described here---doesn't make any assumptions as to what a scaffold should look like. Instead, through exhaustive enumeration, we defer the scaffold recognition task to the user. We feel this approach is reasonable since, after all, even medicinal chemists don't often agree on the definition of "scaffold."
The prototype is available here. Below is a quick look of it.
Since this prototype was put together quickly, its user interface is a bit clunky but hopefully still functional. This is the first time it's been exposed outside of our group, so we'd be very interested in comments and feedback on how we can make it better.
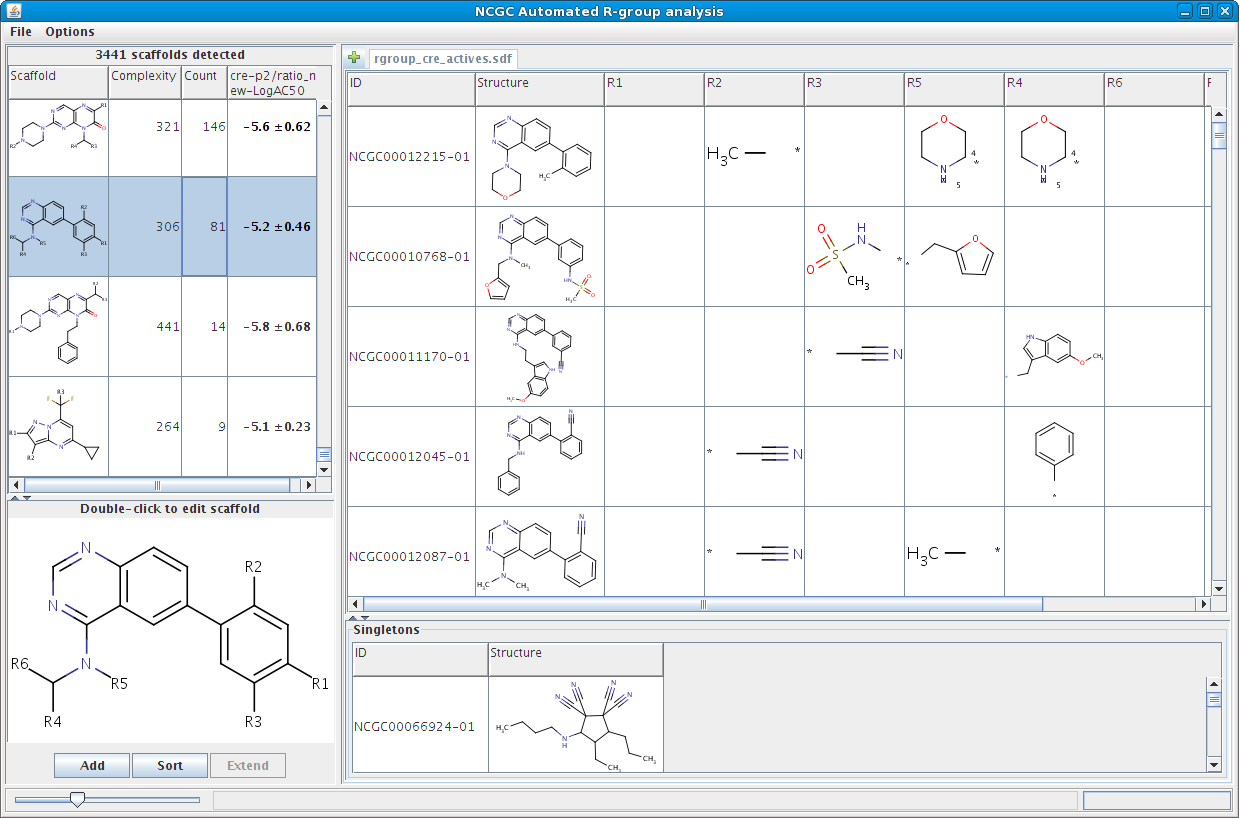
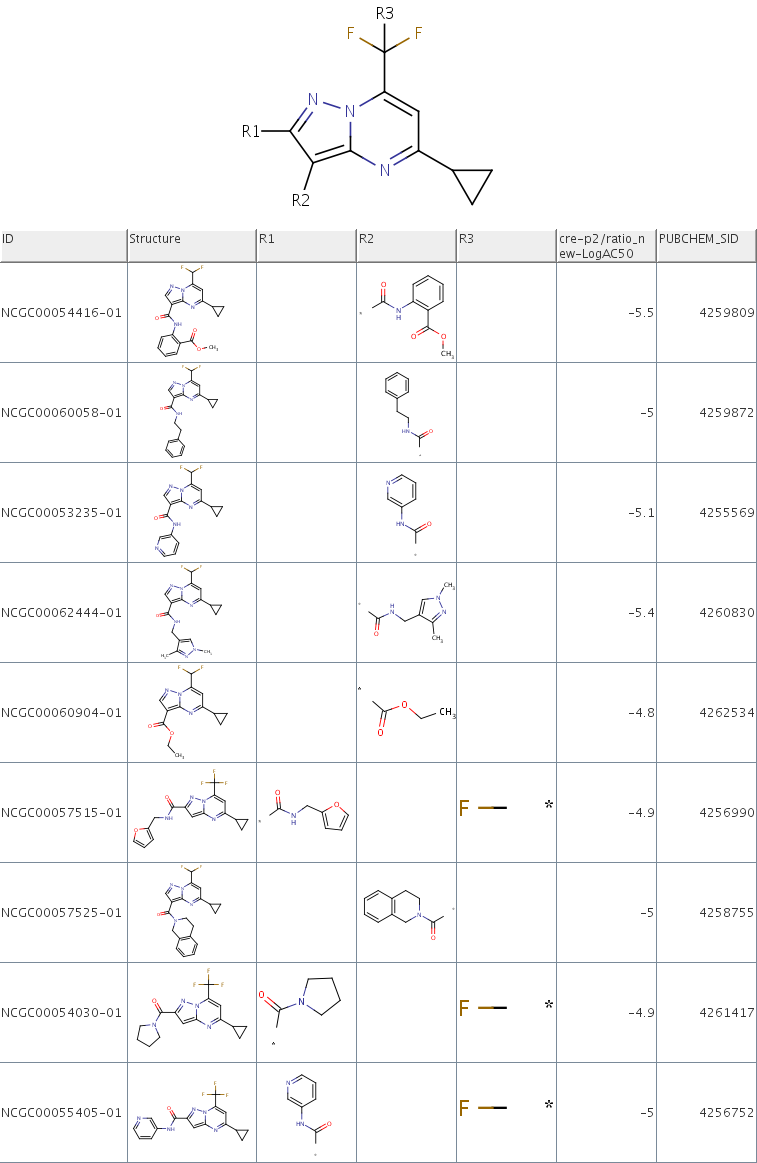

Hi!
I tried to run the program on my MacBook, running OS X 10.6.3. However, after it’s downloaded nothing more happens. It opens up the program but I get no windows or menus or anything. The java console is showing the following:
Reading certificates from 11 http://tripod.nih.gov/ws/rgroup/rgroup.jar | /Users/fwr/Library/Caches/Java/cache/6.0/53/590b2c75-38649bb7.idx
Reading certificates from 342366 http://tripod.nih.gov/ws/rgroup/jchem.jar | /Users/fwr/Library/Caches/Java/cache/6.0/63/64dcf1bf-3375c702.idx
Reading certificates from 111394 http://tripod.nih.gov/ws/rgroup/jide-common.jar | /Users/fwr/Library/Caches/Java/cache/6.0/37/b7b3c25-59845234.idx
Reading certificates from 2942 http://tripod.nih.gov/ws/rgroup/swingworker-0.8.0.jar | /Users/fwr/Library/Caches/Java/cache/6.0/26/2640095a-3f1384df.idx
Exception in thread “AWT-EventQueue-0” java.security.AccessControlException: access denied (java.lang.RuntimePermission accessClassInPackage.apple.laf)
at java.security.AccessControlContext.checkPermission(AccessControlContext.java:323)
at java.security.AccessController.checkPermission(AccessController.java:546)
at java.lang.SecurityManager.checkPermission(SecurityManager.java:532)
at java.lang.SecurityManager.checkPackageAccess(SecurityManager.java:1512)
at sun.misc.Launcher$AppClassLoader.loadClass(Launcher.java:327)
at java.lang.ClassLoader.loadClass(ClassLoader.java:305)
at java.lang.ClassLoader.loadClass(ClassLoader.java:250)
at com.jidesoft.plaf.LookAndFeelFactory.loadLnfClass(Unknown Source)
at com.jidesoft.plaf.LookAndFeelFactory.isAssignableFrom(Unknown Source)
at com.jidesoft.plaf.LookAndFeelFactory.isLnfInUse(Unknown Source)
at com.jidesoft.plaf.LookAndFeelFactory.installJideExtension(Unknown Source)
at com.jidesoft.plaf.LookAndFeelFactory.installJideExtension(Unknown Source)
at com.jidesoft.plaf.LookAndFeelFactory.installJideExtension(Unknown Source)
at com.jidesoft.swing.JideTabbedPane.updateUI(Unknown Source)
at javax.swing.JTabbedPane.(JTabbedPane.java:189)
at com.jidesoft.swing.JideTabbedPane.(Unknown Source)
at com.jidesoft.swing.JideTabbedPane.(Unknown Source)
at gov.nih.ncgc.rgroup.RGroupTool.createContentPane(RGroupTool.java:1000)
at gov.nih.ncgc.rgroup.RGroupTool.initGUI(RGroupTool.java:935)
at gov.nih.ncgc.rgroup.RGroupTool.(RGroupTool.java:867)
at gov.nih.ncgc.rgroup.RGroupTool$JnlpLaunch$1.run(RGroupTool.java:1753)
at java.awt.event.InvocationEvent.dispatch(InvocationEvent.java:209)
at java.awt.EventQueue.dispatchEvent(EventQueue.java:633)
at java.awt.EventDispatchThread.pumpOneEventForFilters(EventDispatchThread.java:296)
at java.awt.EventDispatchThread.pumpEventsForFilter(EventDispatchThread.java:211)
at java.awt.EventDispatchThread.pumpEventsForHierarchy(EventDispatchThread.java:201)
at java.awt.EventDispatchThread.pumpEvents(EventDispatchThread.java:196)
at java.awt.EventDispatchThread.pumpEvents(EventDispatchThread.java:188)
at java.awt.EventDispatchThread.run(EventDispatchThread.java:122)
Any ideas as to what went wrong? I have tried to empty the cache and downloaded it again as well as restarting the computer.
Hi Fredrik, please try again now. You might have to clear your browser’s cache.
Hi again, I just saw your comment and I tried and it’s working…