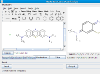
Structure standardizer
Although much of our recent effort has been toward the user interface, the backend of Tripod is by far where we've spent most of our development effort. We hope to provide a more detailed description of the overall architecture of Tripod in a future post, but suffice to say Tripod is a self-hosted application with persistent data storage for biological and chemical entities such as compounds, assays, targets, genes, documents, etc. To make effective use of the entities for downstream analyses (e.g., building polypharmacology networks), the entities themselves must be uniquely registered within Tripod's persistent data store. For the most part this is straightforward, since each of the entity types usually has some form of well-known registry identifier (e.g., UniProt accession for protein targets, locus ID for genes, DOI for publications). The exception, of course, is compound entity. Although there are well-known registries available, using any such registry would severely limit the utility of Tripod, especially within a corporate setting.
Compound registration within Tripod is not quite like a traditional chemical registration system. In addition to assigning unique registry identifiers, Tripod also performs additional processing (e.g., generate fragments and structural indexes) to speed-up downstream analysis such as R-group decomposition. (It's our hopes that any useful analysis task in Tripod will ultimately be reduced to a simple step of browsing.) In terms of structure standardization, Tripod is aggressive at (i) stripping out salt/solvent, (ii) (de-) protonation, and (iii) removal of "spurious" sp2 stereochemistry. Here what we mean by "spurious" is effectively any E/Z configuration that might be induced due to either tautomer/mesomer enumeration and/or alternating path during proton removal (i.e., see Section 5 of the InChI technical manual).
We're now wrapping the standardizer engine used in Tripod's registration system as a self-contained tool in hopes of getting feedback on how it can be improved. While we've done our best to consult InChI and PubChem's standardizer during its development, there are just too many posibilities for us, with limited bandwidth, to have any hope of getting it right (for examples, the different types of tautomeric forms discussed in this paper should provide a good workout for any chemical registration system).
The standardizer tool is available as a Java webstart application here. It requires at least Java 1.5. The tool can be used interactively or in batch mode. Please let us know if you have problems running it. Below is a quick look of it.